Quality DNA from crap samples – The importance of DNA quality in metabarcoding
From detecting probiotic organisms in fermented foods to assessing soil health, microbiome profiling is a critical component of research and development in food, beverage, agricultural, health, and environmental science. Microbiome profiling typically relies on DNA metabarcoding – where a region of the microbe genome that is highly variable between species is used as a means of identification. When a population of microbes are present, amplification of these “barcode” regions and identification by next-generation sequencing enumerates the species present in much the same way that reading the barcodes on grocery items allows identification of the products being purchased. Typically, this barcode region is the internally transcribed spacer (ITS) in fungi and the 16S ribosomal RNA gene (EB16S) in bacteria.
First things first – Extracting Quality DNA
The first step in microbiome profiling using metabarcoding is DNA extraction from the sample. While this might seem simple, it can be anything but. Typical samples such as feces, soil, or fermented foods contain many substances that are difficult to separate from DNA, and which strongly inhibit downstream steps such as PCR. Although many DNA extraction kits can do a decent job of purification, they are no substitute for experience and knowledge when it comes to getting top quality DNA from diverse sample types. Often kits fail at the first step – extraction. Having the right equipment and procedures for sample homogenization are critical for ensuring that DNA extracted from samples is representative of all the organisms present. At Proxima, we use proprietary protocols designed to maximize extraction and purity of DNA from specific sample types to ensure accurate and representative microbiome profiling.

Assessing DNA Quality and Yield
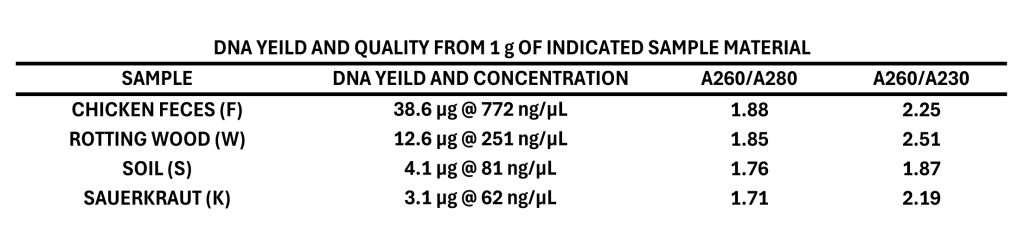
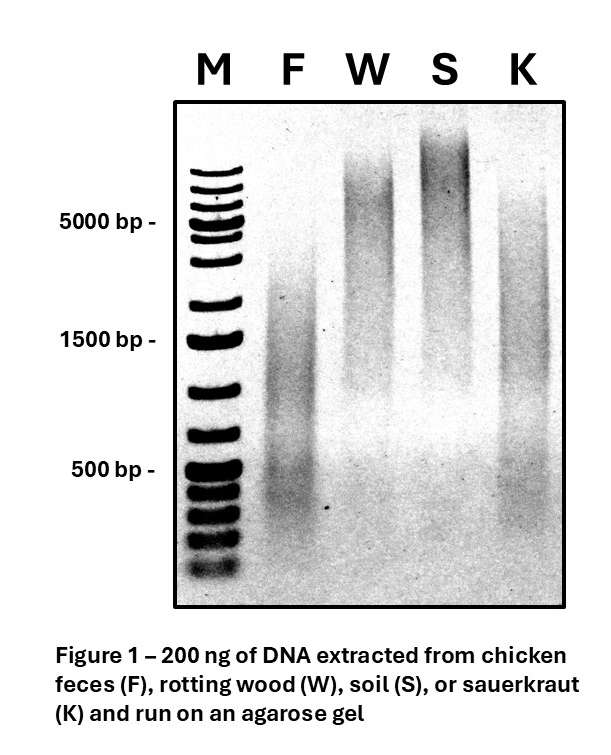
Figure 1 shows 200 ng of DNA extracted from four different types of samples which are typically considered difficult to obtain quality DNA from, and the attached table shows the yield and quality of DNA extracted from one gram of each of these sample materials. Samples used were chicken feces (F) aimed at analyzing differences in gut microbiome between free-range and traditionally raised birds; rotting wood (W) from the boreal forest with an aim of identifying organisms producing cellulases and other enzymes with applications in biofuel production; soil (S) looking for the presence of fungal diseases and assessing soil health; and a naturally fermented sauerkraut (K) with the aim of studying the range of probiotic organisms present. A key indicator of quality DNA for metabarcoding is the presence of significant DNA above 800 bp in length, which is required for efficient amplification of the ITS and EB16S sequences. DNA should also have an A260/A280 ratio between 1.70 and 1.90. The data shown here demonstrates how Proxima can efficiently and effectively extract high purity DNA from difficult samples for barcoding applications.
Amplifying DNA Barcode Regions with PCR
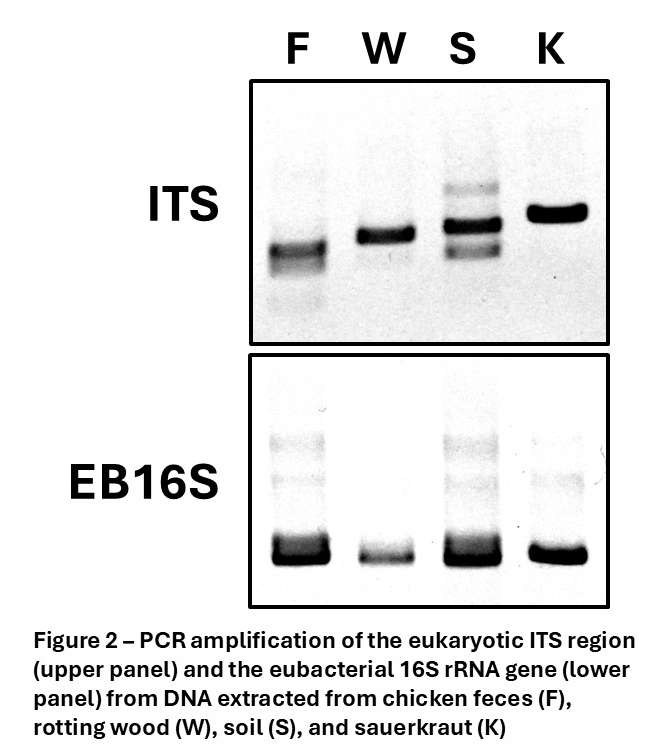
The next step in microbiome profiling is PCR amplification of the barcode region, whether ITS for analysis of fungal content or the eubacterial 16S rRNA gene for bacterial microbiome analysis. The most common issue with DNA extracted from samples such as those shown here is the presence of PCR inhibitors which prevent efficient amplification of barcode regions. In Figure 2 we show the results of PCR reactions employing the purified DNA shown above and standard ITS or EB16S primers. PCR was efficient and high yielding in all samples, showing the absence of PCR inhibitors and ensuring an accurate representation of organisms present.
Result – Successful Microbiome Profiling
Whenever and however microbiome profiling fits into your research, trust Proxima to ensure accurate results from any sample type at affordable prices and industry leading turnaround. Book a free consultation to discuss your project. At Proxima, we love to talk science.