Masquerading molds – The importance of genotyping for fungal identification
We all know that we can’t trust everything we see online. But fungi have been faking their appearance since long before the internet. Microorganisms, and other living things, can be identified and characterized based on two attributes, their genotype and their phenotype. When reporting a phenotypic description, we are reporting observable characteristics of the organism such as their appearance and behavior. Unfortunately, these traits can be ambiguous and difficult to differentiate even to a trained eye, especially when it comes to fungi.
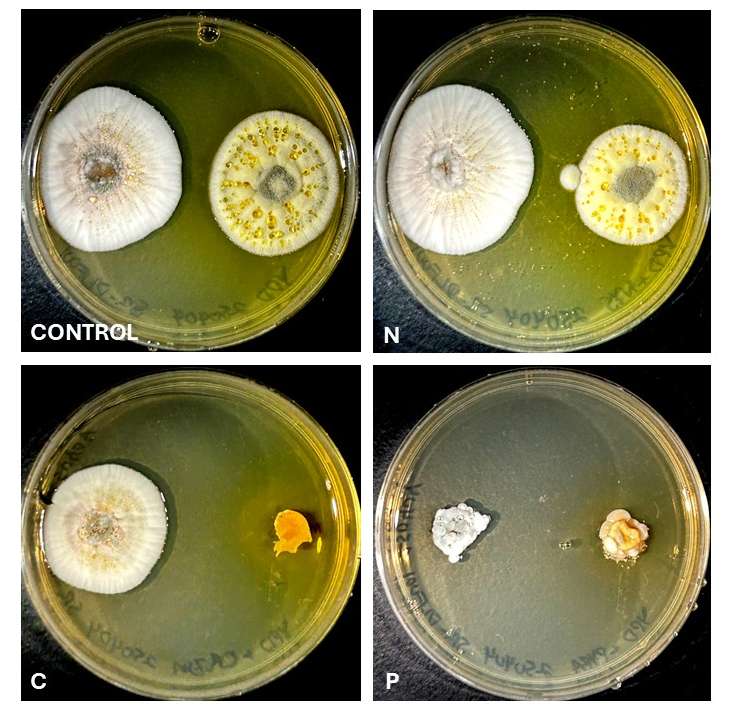
Look at the photos of four fungi growing in agar plates and see if you can guess which organisms are the same species and see how many you got right!
Things are not Always What they Seem
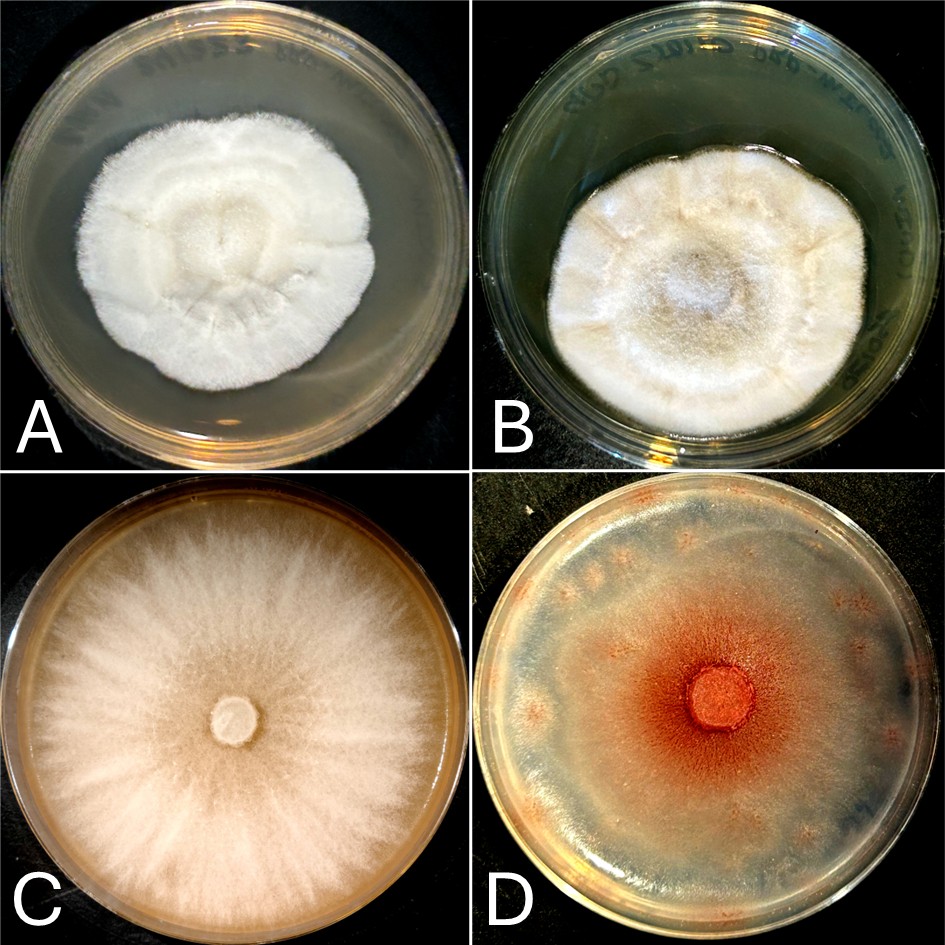
If you thought (A) and (B) were the same species, you’d be wrong. Isolate (A) is a Neoascochyta spp. and (B) is Clonostachys spp. If you thought (C) and (D) were different species, you’d be wrong again. They are the same species and isolate of Fusarium growing on two different media compositions, however, only the medium in (D) contains the necessary components to produce pigment! This demonstrates how an organism’s appearance, the phenotype, can be affected by different environmental conditions and available nutrients.
Traditionally, fungi would be identified based upon differences in growth on differential media and microscopic examination of spore-bearing structures and other features. However, aside from the considerable time and effort involved, this process can be inaccurate as certain structures may not always be present in each species under a given set of growth conditions, as the figures show. This is why identification of fungi through a process termed “genotyping” is incredibly important in the laboratory setting, providing greater accuracy with less investment of time and materials.
DNA Barcoding for Fungal Identification
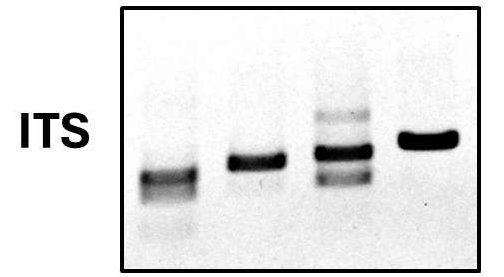
Identification of fungi by genotyping is accomplished via PCR amplification and sequencing of a region of the genome called the internal transcribed spacer sequence or ITS. The ITS is a DNA sequence found between two regions that encode the ribosomal 18S and 5.8S RNAs. Since this spacer region does not code for any functional cellular components it is prone to accumulating point mutations over the course of evolution. Due to this, the sequence of the ITS is highly variable between different species and can be used as a means of species identification.
In other words, the ITS can be sequenced to determine the species in a similar way to a barcode being read in the grocery checkout to identify the product being purchased. This similarity gives rise to the use of the term “DNA barcoding” to mean the use of a small highly variable DNA sequence for species identification. Whether dealing with common pathogens like Fusarium or exotic fungi like Neoascochyta, at Proxima we use DNA barcoding to quickly and efficiently ensure we achieve accurate fungal identification when it matters most.